Researchers are often interested in using a pooled shRNA library for genome-wide RNAi screening to cast a very “wide and unbiased net to identify any and all genes functionally involved in some pathway”. Although it is not difficult to make an shRNA library targeting all human or mouse genes, it is practically very difficult to comprehensively screen using such a library. Careful consideration of starting cell numbers and handling of cells during propagation is essential to ensure thorough screening of pooled shRNA expression libraries, minimize false negatives, and obtain consistent and reproducible results.
First, there is an issue of library complexity since it is necessary to have several shRNAs designed to target each gene. The effectiveness of “validated” shRNA varies from cell-to-cell, and no effective shRNA has been identified for many genes. For these reasons, it is necessary to incorporate several shRNAs for each gene to ensure reasonable knockdown of a high percentage of targets. Cellecta typically designs 5-6 shRNA against each target gene, so more than 25,000 shRNA are required to target 5,000 genes. A library targeting the entire human genome, estimated at just over 23,000 genes, requires approximately 115,000 individual shRNA constructs. While it is not particularly difficult to construct libraries of this complexity, this number of unique shRNA sequences creates technical challenges with representative screening.
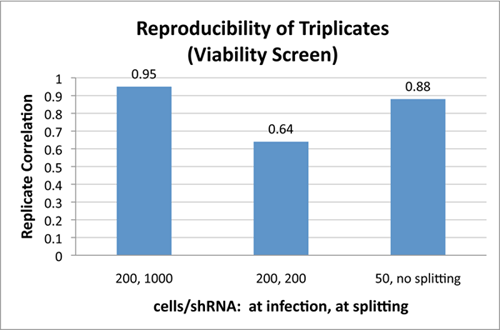
Pooled shRNA library screens require quantification of changes in the fraction of each shRNA sequence in selected vs. control cells or starting library. A “hit” occurs when selected cells have significantly more or less of a particular shRNA sequence. Whether one is looking at enrichment of specific shRNA in the selected cells vs. the control (positive selection) or depletion of shRNA in selected cells vs. the control (negative selection), it is critical that the screen begin with sufficient numbers of each shRNA to ensure measured changes in the fraction an shRNA sequence are statistically significant. This means that, if there are very low numbers of specific shRNAs at the start of the screen, small random changes in a drifting population may be difficult to differentiate from significant trends. Simply put, a loss of 2 shRNA is a 20% change if there are only 10 initially vs. 2% if there are 100. For this reason, a least a few hundred cells need to be infected with each shRNA to initiate a good screening. This is demonstrated in the data below where starting with a smaller population of just 50 cells per shRNA (third bar) leads to more variance than starting with a population of 200 cells per shRNA (first bar). This means that starting a screen involves infecting 100 times more cells than the complexity of the library. For a library with 25,000 shRNAs, the starting population should be 2.5 million infected cells, and for a library with 115,000 shRNAs, the starting population should be over 11 million infected cells.
To screen a heterogenous mixture of shRNA expression constructs, however, it is important to have 2-3 times more cells than viral particles to help ensure that most cells are only infected with one shRNA-carrying virus (i.e., a multiplicity of infection [MOI] of 0.3-0.5), so you need to have 2-3 times more cells than the number targeted for infection. Thus, 6-8 million cells are needed to start a screen with libraries of 25,000 shRNAs, and a whole genome library of 115,000 shRNAs would require 25-35 million cells. Since each screen should be done in duplicate, or better, triplicate, the number of cells needed makes a full genome screen with a redundant shRNA library impractical.
Finally, to ensure a comprehensive screen, it is not simply sufficient to start with the right amount of cells. During the screening process, incorrect propagating the cells can completely undercut the representation set up at the initiation of the screen. This is especially true for a negative selection screen, such as a viability screen where one is interested in identifying shRNA that kill or inhibit proliferation of cells, and, therefore, drop out of the population. It is critical to maintain the full library representation that was initially used at the start of the screen. If a portion of propagating cells are removed during propagation (e.g., cells are split), the representation of the library can be skewed in the sample. By doing so, this introduces significant noise. This effect is readily seen in the first two bars of the figure where the effect of starting with sufficient cells (200 cells per shRNA) is completely undercut by splitting cells during propagation so that that the final count of cells after 10 days is the same as the initial number of cells. The correlation between triplicates falls dramatically when the cells are split.
Library representation is often overlooked, especially when the desire is for large-scale unbiased screens. However, without careful consideration in designing screening procedures that reflect the complexity of the library, results of these large-scale screens can produce relatively meaningless data with anecdotal results at best. So, what about genome-wide screening? Our approach with the DECIPHER Project is to provide modules, each targeting approximately 5,000 genes with 27,500 shRNA, which enable comprehensive screening. We are more than half way finished building a series of 5 modules that will target all human genes, and 4 modules targeting all mouse genes. For more information on these libraries, visit the DECIPHER Project website.


